Metabolomics in the Context of Forensic Omics Biomarkers
LS Castillo-Peinado1,2, MD Luque de Castro1,2*
1 Department of Analytical Chemistry, Annex Marie Curie Building, Campus of Rabanales, University of Córdoba, Córdoba, Spain.
2 Maimónides Institute of Biomedical Research (IMIBIC), Reina Sofía University Hospital, University of Córdoba, Córdoba, Spain.
*Corresponding Author
M.D. Luque de Castro,
Maimónides Institute of Biomedical Research (IMIBIC),
Reina Sofía University Hospital, University of Córdoba, E-14071, Córdoba, Spain.
Department of Analytical Chemistry, Annex Marie Curie Building,
Campus of Rabanales, University of Córdoba, Córdoba, Spain.
Email: qa1lucam@uco.es
Received: October 30, 2017; Accepted: November 27, 2017; Published: November 30, 2017
Citation: LS Castillo-Peinado, MD Luque de Castro. Metabolomics in the Context of Forensic Omics Biomarkers. Int J Forensic Sci Pathol. 2017;5(7):384-391. doi: dx.doi.org/10.19070/2332-287X-1700082
Copyright: MD. Luque de Castro© 2017. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.
Abstract
Search for biomarkers is supported on any of the great omics, but the given omics is rarely discussed or even named in the literature as being the basis of the target biomarker. In this context, metabolomics biomarkers exist even before this omics discipline was defined as such, but the fact that a biomarker is based on the changes in the concentration of one or several metabolites does not seem to be enough reason for giving to it the name of metabolomics biomarker. This omission is discussed here for forensic omics in general and for forensic metabolomics in particular. Present forensic biomarkers belonging to different omics are compared and the present and foreseeable future of forensic metabolomics biomarkers discussed.
2.Abbreviations
3.Introduction
3.1 Definition of Biomarkers and their Role in Forensics
3.2 Single and Multiple Biomarkers
3.3 Scope of Forensics Biomarkers
3.4 Pursued Objectives
4.Omics in the Context of Forensic Biomarkers
5.Analytical Tools for Search and Determination of Biomarkers in Forensic Metabolomics
6.Present Metabolomics Biomarkers in Forensics
6.1 Metabolomics Biomarkers that Surpass those based on other Omics: Alcohol Biomarkers
7.Future of Metabolomics Biomarkers in Forensics
8.Acknowledgements
9.References
Keywords
Biomarkers; Omics Disciplines; Forensic Metabolomics.
Abbreviations
AMI: Acute Myocardial Infarction; BAC: Blood Alcohol Concentration; CDT: Carbohydrate deficient transferrin; CF: Cystic Fibrosis; CK: Creatine Kinase; CODIS: Combined DNA Index System; CRP: C-Reactive Protein; DSM: Mental Disorders; EtG: Ethyl Glucuronide; EtS: Ethyl Sulfate; LC–MS/MS: Liquid Chromatography Coupled to tandem Mass Spectrometry; LC–MS QqQ: Liquid Chromatography Coupled to Mass Spectrometry with Triple Quadrupole; LDH: Lactate Dehydrogenase; LISH: Ligation in situ Hybridization; MALDI: Matrix-Assisted Laser Desorption/ Ionization; MS: Mass Spectrometry; PEth: Phosphatidylethanol; PK: Pyruvate Kinase; PSA: Prostate-SpecificAntigen; Q:Quadrupole; QTOF: Quadrupole Time-of-Flight; UHPLC: Ultra-High Pressure Liquid Chromatography.
Introduction
Scientific investigations on biological traces occurring on a crime scene are carried out through a series of sequential steps involving generic diagnosis, species diagnosis, regional diagnosis and individual diagnosis. All together these steps allow researchers to reconstruct the dynamics of the criminal event and to check the reliability of assertions by putative suspect people. The so-called biomarkers help in the successful development of one, several or all the steps in the series.
The term ‘biomarker’ has been defined differently depending on the scientific area to which it belongs. For example, in toxicology (also applicable in the forensics area) a biomarker could be defined as ‘a component (or components) in biological fluids, cells, tissues, or whole organisms that indicate the presence, magnitude, and exposure of toxicants or of host response’ [1]. In the clinical area (also applicable in forensics) a biomarker is defined as ‘a characteristic that can be objectively measured and evaluated as an indicator of a physiological orpathological process or as a pharmacological response to a therapeutic intervention’ [2]. Otherwise, the World Health Organization defines a biomarker as ‘any substance, structure, process or its products that can be measured in the body and influence or predict the incidence of outcome or disease’. In spite of all the different definitions of biomarker, they all coincide on the idea of a biomarker as almost any measurement reflecting an interaction between a biological system and a potential hazard, which may be chemical, physical, or biological [3].
Even though biomarkers are by definition objective and quantifiable characteristics of biological processes, they may not be necessarily associated to an individual’s experience or health status. This is the main reason why biomarkers identification requires their relevance and validity to be established. Thus, their assessment involves repeated evaluation taking into account distinct points of view in order to establish its reliability to characterize a specific situation or the individual health status [3]. Final validation is based on comprehensive application to a number of cases that ensures representativeness.
In forensic sciences, the term ‘biomarker’ has different meanings depending on the forensic discipline and some definitions coincide with those used in other areas, as is the case with toxicological or clinical biomarkers. In dealing with forensic environmental geochemistry, biomarkers are defined as compounds derived from biological sources (i.e., natural products) that retain some, if not all, of the structural characteristics of their parent precursor molecule after being preserved in the geological record or released into the environment [4]. Even though the use of biomarkers in forensic science still needs notable research, it is highly spread through the different areas in this field, as can be found in the literature [5, 7].
Biomarkers can be classified attending to different criteria; one of them is the capability to characterize an organism situation (single biomarker) or the necessity of other biomarkers for complete characterization (multiple biomarkers). An example is found in biomarkers of alcohol consumption, which can reveal recent alcohol exposure or a prolonged risk of alcohol consumption. Among the latter are carbohydrate deficient transferrin, γ- glutamyltranspeptidase and phosphatidylethanol. The last is present in the body only after alcohol intake, while the first two are influenced by factors different from alcohol intake such as sex, age and smoking habit, among others [8]. Therefore, in order to obtain a more complete and unequivocal information multiple biomarkers should be used. A recent and interesting study on the dynamics of multiple biomarkers in mouse excisional wounds for wound age estimation (from 12 h to 10 days) was based on neutrophil and macrophage infiltration, fibroblast and fibrocyte accumulation, their myofibroblastic transformation and some wound healing-related molecules [9]. One other classification of biomarkers is based on the number of compounds forming part of the given biomarker. This classification is characteristic of metabolomics biomarkers, giving place to single metabolite biomarkers and panel metabolite biomarkers [10, 11].
The sequential steps to which forensic scientists face in their research involve: first, to determine whether a crime has been committed; if so, their assistance focuses on identification of the offender; then, to conduct detailed studies in different areas depending on the evidences they have at their disposal [12]. The areas are specific and well differentiated; this is the main reason why biomarkers are not considered in the same way when scientists work with clinical, environmental or geological samples, among others. Despite forensic biomarkers for clinical samples have so far been more abundant than for environmental samples, advanced techniques and databases have recently been developed in environmental forensic studies to investigate the source, transport, and fate of organic compounds present in the environment. Biomarkers in this discipline consist of compounds that are contained in the target environmental sample (oil, soil, air, etc.), and characterize, correlate and differentiate the sample from others. In this way the source of identification, the determination of its certain composition and distinct relievable factors for the crime investigation are allowed. Nowadays, several spillages of refined oils frequently occur during their production, transportation, storage, and use. These may cause significant environmental problems and this is the reason why researchers perform fingerprinting analysis of oil spills for site contamination assessment, and they also determine the legal liability for the spill [5]. These biomarkers are one of the most important hydrocarbon groups in petroleum for chemical finger printing, which are usually stable and show little or no changes in structures from their parent organic molecules [6]. However, recent research has demonstrated that different factors, such as weather, could be involved in the changes [7].
In dealing with clinical samples, forensic biomarkers are particularly desirable. These biomarkers are very helpful, for instance, to evaluate elder abuse - including abrasions, lacerations, bruising, fractures, restraints, decubiti, weight loss, dehydration, burns, cognitive and mental health problems, hygiene issues, and sexual abuse [13]. Their identification is crucial to the medical and legal establishment of whether elder abuse or neglect has occurred [14].
The aim of this article was first to give a call of attention to the fact that biomarkers in general and forensic biomarkers in particular are supported on given omics that must be specified by the authors; then, to discuss the present and foreseeable future of forensic metabolomics biomarkers.
Omics in the Context of Forensic Biomarkers
Since Jeffreys et al. introduced in 1985 the DNA fingerprinting in the field of forensic genetics, proving that some regions from the DNA contain repetitive sequences which are variable among individuals [15], the results of DNA analysis have been accepted as evidence in the court in many countries [16]. When in 1992 the European Council issued the Recommendation No. 92, regarding the use of the DNA analysis in the criminal justice, the DNA test was accepted in the Court [17].
Forensic genetics has experienced a strong development during the last 20 years supported on the new discoveries and technologies in molecular biology. Identification of people present in a crime scene is based on analysis of DNA samples extracted from biological fluids such as blood, saliva, semen, or urine; or from tissues such as bones or hairs. DNA analysis has long been able to provide information about an individual’s biological sex [18, 19]. More recently, it has also been shown that DNA can be used to learn other externally visible characteristics about the donor, such as eye color, race, and sex [20]. DNA analysis is strongly considered the gold standard in forensic analysis of blood samples because of its highly accurate results (if a DNA match is made the chance of it being a false positive or a coincidental match is on the scale of 1 in 34 million [21]). This is due to the comparative assessment between the genetic profiles obtained from biological traces found on the crime scene and the profile of a suspect. The profile is entered and logged into the Combined DNA Index System (CODIS) so that a comparison can be made to a person who has been previously arrested [15, 22, 23]. If there is no suspect in custody or no match in the CODIS database either the chromosomal DNA found in the nucleus or the mitochondrial DNA, the profile is essentially useless for the police. Other no minor shortcomings of DNA analysis are the time, price, and technical skill required to analyze a single blood sample.
Continuous improvements in sample preparation in genetics have a clear example in the evolution experienced from the known as Nano String technique to handle damaged mitochondrial RNA which requires both expensive instrumentation and reagents and has limited sensitivity to ligation in situ hybridization (LISH), cheaper and less expensive. Addition of LISH probes to damaged mitochondrial RNA allows turning them into sequences of mostly DNA that can be amplified and sequenced without reverse transcription [24].
Proteins, contrary to DNA, were rarely considered as sources of useful biological traces during the first years of proteomics life and were scarcely used in crime scene investigations because: (i) they tend to be less stable than DNA and are easily degraded. (ii) Individual diagnosis by protein identification is impractical because protein variability is low. (iii) The amount of sample available cannot be amplified by “PCR-like” procedures--this aspect can be partially overcome by the possibility of including the original biomarker in a single biocatalytic cascade for amplifying its effect on the final analytical output signal [25]. (iv) Generally, higher amounts than in genomics or extremely high sensitive analytical techniques are required in proteomics. After more than 20 years of proteomics research [26], also the forensic area has been highly benefited from this omics. For example, proteomics has proved that the biological sex, ethnicity, and age of a blood sample can all be determined using multi-enzyme assays [27, 28]. This type of system was used to conduct research using the markers creatine kinase (CK) and lactate dehydrogenase (LDH), in combination with pyruvate kinase (PK) to distinguish between African American and Caucasian blood sample originators [25]. In a similar reaction combining CK, PK, LDH, and alanine transaminase sex of a blood sample originator was determined [28, 29]. In analyzing the use of this assay the samples were allowed to age for up to 120 h and it was determined that the assay could also be used to know the time since a sample was deposited [30]. In addition, a oneenzyme biocatalytic cascade has been used to distinguish between young and old people, independently of the sex, by analysis of blood alkaline phosphatase based on photometric monitoring of p-nitrophenol at 450 nm [31]. In this way, the traditional methods applied in forensic serology (usually based on immunoassay [32] or DNA [33]/RNA [34] analysis) with involvement of sophisticated equipment and highly skilled personnel [35] can be substituted by a simple enzymatic assay performed directly at a crime scene, thus giving rapid results for identification of a pool of possible suspects. Given the value of modern crime scene DNA analysis for gathering specific information on the age, sex and race, the enzymatic assay cannot substitute for a DNA ‘fingerprint’ as it only narrows a group of suspects with some uncertainty. However, its advantage lies in quick, on-site analysis, which is markedly absent in modern crime scene analysis.
The development of a proteomic-based approach to identify unambiguous protein biomarkers for each individual matrix with the aim to constitute a specific database for matrix characterization fulfilled the requirements of forensic science for the rapid determination of biological matrices. In fact, mass spectrometry (MS)- based proteomics procedures can be used to identify the most prominent proteins specifically present in each biological matrix found at a crime scene (blood, saliva, semen, vaginal fluid, nasal secretion or urine). One single test is sufficient to determine unambiguously the identity of the biological matrices, even in complex mixtures. In addition, species identification is also possible, and the analysis is performed on the first “washing” step of DNA extraction, thus saving most of the sample for subsequent DNA analysis. The protein biomarkers for the most frequently encountered biological matrices (saliva, blood, semen and lacrimal fluid) have been clearly identified in real forensic samples [31].
Concerning metabolomics - the youngest of the great omics, the field it and its subdisciplines open to the forensic science is in developing yet. Isolated incursions in metabolomics subdisciplines for forensic biomarkers were made years ago. Boccardet al., in 2011, proposed an approach based on an analytical platform involving an ultra-high pressure liquid chromatograph (UHPLC) and a quadrupole time-of-flight (QTOF) mass spectrometer for untargeted steroidomics in urine samples from a clinical trial for the discovery of relevant biomarkers of testosterone undecanoate oral intake. In this way, a wide door was open to the untargeted simultaneous evaluation of a high number of potential biomarker candidates [36]. Since then, metabolomics has provided numerous biomarkers in the clinical area in general and in forensics in particular, despite it is hardly taken into account most times. For example, in the characterization of the salivary epigenome, the importance of previous work in dealing with the salivary microbiome, proteome, endocrine analytes, genome, and transcriptome was highlighted [37], but the possible role of metabolomics was
ignored, despite its crucial importance in unraveling epigenetic behavior [10]. At present, the importance of metabolomics biomarkers increases at a fast rhythm [38,39], even surpassing wellestablished proteomics biomarkers. This is the case with ethanol consumption in which metabolites from this drug are clearly better biomarkers of chronic drinkers than well know indirect biomarkers such as enzymes (e.g., aspartate transferase, alanine transferase, γ-glutamyltransferase), mean corpuscular volume of the erythrocytes or carbohydrate-deficient-transferrin, as can be seen in a section below.
It is remarkable that, despite an omics study precedes the discovering of a new biomarker, most authors do not specify in their publications the omics to which their investigated biomarkers belong, but only the sample and their forensic use [40].
Analytical Tools for Search and Determination of Biomarkers in Forensic Metabolomics
Traditional assays using as detector the naked eye after spraying on the target common reagents such as phenolphthalein or luminol, or even using ultraviolet light, for in situ analysis in the crime scene, the most popular and the only known by people addicted to television police series, are not the subject of this section. We consider here equipment for analysis of metabolomics biomarkers proposed in the literature and its desirable evolution in the light of the present and foreseeable developments.
Distinction is made between the search for metabolomics biomarkers and their quantitation in a given sample once the panel has been assessed; that is, between untargeted and targeted metabolomics analysis, respectively.
The first and crucial step in the search for metabolomics biomarkers or for their quantitation is sample preparation. This step can vary from direct insertion of the sample either into the detector (or more commonly into previous separation equipment), to the use of microfluidic (chip-based) devices coupled to capillary high-resolution separation tools [41], or solid-phase extraction commercial equipment on-line connected to LC-MS triple quadrupole (QqQ)arrangements [42, 43]. These devices constitute a milestone in forensics equipment for analysis of small samples.
Equipment based on time-of-flight (TOF)-either with quadrupole or matrix-assisted laser desorption/ionization (Q or MALDI, respectively)--coupled to liquid or gas chromatographs, depending on the features of the metabolites, is commonly used for identification, followed by metabolites selection using chemometric approaches (ANOVA, PCA, PLS-DA, heat maps, ROC curves, etc.) [38, 39, 42]. The sensitivity and specificity displayed by each metabolite in the ROC curve clearly establish its usefulness as biomarker.
The preferred option for targeted analysis of metabolomics biomarkers is MS in the QqQ approach, as it provides one of the most sensitive tools for metabolites quantitation [10, 43, 44]. It should not be forgotten that absolute quantitation by this tool requires preparing standard curves in the same matrix as the samples under study so that the matrix effect and the extraction efficiency for the target metabolites must be the same in both the standards and samples. Taking into account that, by definition, analyte-free biological matrices do not exist for endogenous compounds, the lack of blank matrices for quantification of endogenous compounds by LC-MS/MS must be addressed by different approaches (e.g., standard addition, background subtraction, surrogate matrix, and surrogate metabolite methods) by comparing their advantages and disadvantages [44].
A desirable option in metabolomics biomarkers is the use of no destructive analysis, particularly when the sample is scant and several parameters should be analyzed in it. A discussion on the application of no destructive techniques in metabolomics biomarkers analysis is given in the section below.
The trend to miniaturization of analytical instruments leads to portable equipment either for identification or quantitation of metabolomics biomarkers for their on site use. Fast and accurate analysis, as facilitated by this equipment, could be crucial in some forensics research. An example is the Huynh et al.,’s proposal for the use of a portable Raman instrument for analysis to be done entirely on site, thus avoiding any risk of sample damage [45].
Present Metabolomics Biomarkers in Forensics
To omit the name of the omics to which the biomarker belongs is a general situation in forensics biomarkers, more remarkable in the case of metabolomics biomarkers. It could be a consequence of the use of metabolites as biomarkers before metabolomics was defined, as the determination of chloride in sweat as biomarker of cystic fibrosis (CF) [46]. An enormous number of metabolomics biomarkers are daily used in forensics analyses without mentioning this omics (even without awareness of the existence of metabolomics by the forensic analyst). Most of the published articles on the proposal of new metabolomics biomarkers ignore the omics to which they belong. A representative example is the article by Halámek et al. on the sex determination by metabolites (amino acids in this case that they recognize as metabolites) in the sweat contained in fingerprints [45]. This sample has also been used previously to discriminate people from different demographic groups and sexes [47], who had also been discriminated using more invasive sampling as blood [27]. Even age has been determined from blood found at a crime scene [30] without mentioning metabolomics as the discipline whose previous development has allowed the target metabolite(s) to be established as biomarkers.
Countless are also metabolomics biomarkers of drug intake (including alcohol), clearly based on the determination of metabolites from the drug found in samples as varied as hair [40], blood [8], urine [36], saliva [48], human breath [49] or sweat [50].
When biomarkers of a given target situation based on different omics exist, that (or those) based on metabolomics usually offer advantages as compared with those based on other omics.
The fact that metabolomics biomarkers clearly surpass those based on any other omics with similar aims is supported on the requirement by the formers of cheaper reagents, simpler sample preparation and shorter analysis time. A representative example are ethanol biomarkers, as this is one of the most analyzed drugs in a daily basis in forensic and clinical cases of chronic and sporadic alcohol misuse, including driver licensing, workplace testing, and abuse rehabilitation. Both indirect and direct biomarkers of ethanol intake have been widely studied using blood as preferred sample in the former case and a variety of samples (blood, hair, urine, etc.) in the latter [51].
Indirect biomarkers are used in Europe for excessive alcohol intake screening. However, there is not standard protocol for selecting the most appropriate biomarkers to determine alcohol consumption or a defined cut-off level. Furthermore, in North America, indirect parameters of ethanol consumption are not systematically applicable according to the law [52]. This situation is a consequence of the low specificity and sensitivity of indirect biomarkers, as commented in the introduction section [8], since they can be affected by factors different from alcohol consumption. Examples of indirect biomarkers are carbohydrate deficient transferrin, γ-glutamyl transpeptidase, aspartate transferase, alanine transferase, mean corpuscular volume of the erythrocytes, and even serotonin metabolites such as 5-hydroxytryptophol, which allow consumption detection in body fluids (mainly in blood) providing different type of information.
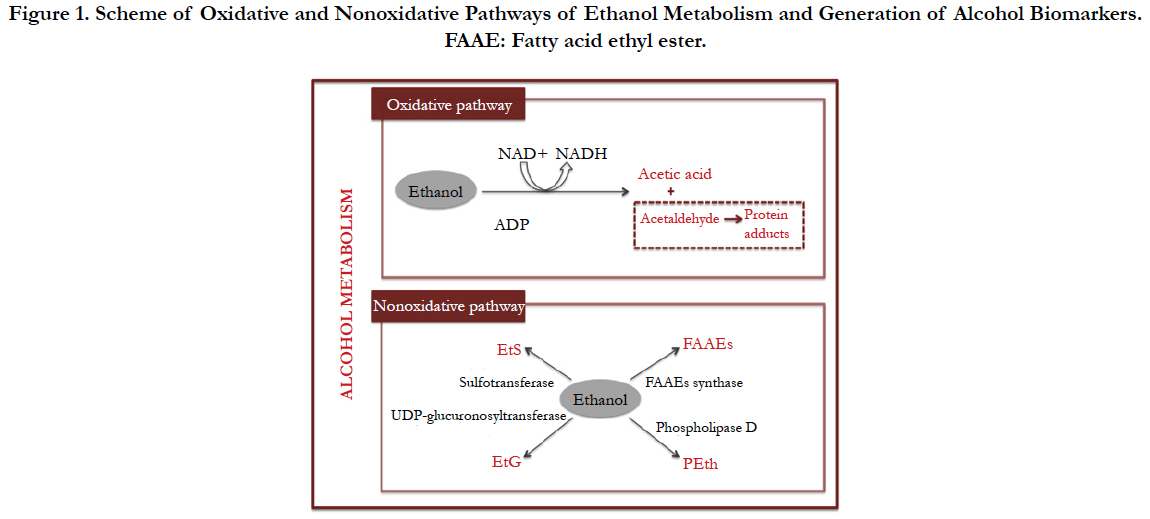
Direct biomarkers of alcohol consumption --e.g., blood alcohol concentration (BAC), ethyl glucuronide (EtG), ethyl sulfate (EtS), phosphatidylethanol (PEth) - have also been widely investigated [8]. Figure 1 schematizes the oxidative and no oxidative pathways of ethanol metabolism and formation of the biomarkers from it [51]. Particularly EtG proves to be very sensitive and specific for identification of active heavy drinkers, and guarantees excellent diagnostic accuracy-with negligible false positive and false negative results, provided an appropriate cutoff value is established [52]. Owing to its short half-life in blood, EtG is not usually determined in this biofluid; therefore, EtG in urine is more commonly used, while hair is used as a longer marker of ethanol use [50]. Fatty acid ethyl esters also increase with chronic alcohol consumption, and are deposited in hair with comparable mechanism and sensitivity/specificity to EtG [53].
Figure 1. Scheme of Oxidative and Nonoxidative Pathways of Ethanol Metabolism and Generation of Alcohol Biomarkers. FAAE: Fatty acid ethyl ester.
PEth is a direct alcohol metabolite present in cell membranes with a half-life of four days [54]. Up to 48 different PEth homologues have been identified and demonstrated their suitability in forensics analysis as biomarkers of alcohol consumption by a simple, easy and cheap test. The selection of one or other of the homologues depends on how old ethanol consumption is searched, allowing, in the case of PEth, differentiating between social drinking behavior (PEth<700 ng/mL) and excessive alcohol consumption (PEth≥700 ng/mL) [8]. The combination of analyses of biomarkers related to alcohol intake in different samples is very useful to provide predictive models for the diagnosis of alcohol misuse with forensic purposes. For instance, recent studies deal with indirect biomarkers in blood and direct ones in hair [51]; thus is, use of multiple biomarkers.
Future of Metabolomics Biomarkers in Forensics
The search for new metabolomics biomarkers that fulfill the necessities of forensic sciences should encompass those considered below.
Biomarkers for which counterparts based on other omics exist, but that can be surpassed by new metabolomics biomarkers because these can be simpler, cheaper, with shorter response time, etc. Examples should be metabolomics biomarkers of vaginal exposure to semen, which overcome the shortcomings of present biomarkers based on other omics such as those of seminal plasma, among which prostate-specific antigen (PSA) is the best characterized; and biomarkers of spermatozoa and other cells present in semen; none of which are ideal biomarkers [55-57]. The useful and interesting substitution of PSA by a metabolites panel as that recently reported for prostate cancer biomarkers [11] could be an example for promoting forensic research in this area.
Traditional metabolomics biomarkers that can be improved either by changing the type of sample in which they are investigated or by changing the target metabolite biomarker using the same sample. Despite increased levels of chloride in sweat is a well established metabolomics biomarker for CF, present studies consider saliva of CF patients contains increased calcium (resulting in insoluble calcium–protein complexes) which causes turbidity of saliva [58, 59], elevated phosphate levels that could explain a higher occurrence of calculus [60], and more neutral lipids, phospholipids and glycolipids as a consequence of the altered physico–chemical properties of saliva [61]. In addition, there are elevated levels of urea, uric acid, and total protein, especially in submandibular saliva [62]. A multi- or single-metabolite biomarker could easily monitor CF in saliva, which offers a simpler, faster, and cheaper sampling than sweat by the commercial MacroductR sampler.
Metabolomic biomarkers proposed in the clinical area that can be useful in forensic analysis. New biomarkers for cardiovascular diseases such as acute myocardial infarction (AMI) based on gene expression analysis [63] or serum-based proteomics have been investigated [64] for a diagnostic better than that provided by traditionally used biomarkers (e.g., ST-elevation or saddle back pattern of Q-wave [65]). The search of the authors’ group for evaluation of coronary artery disease constitutes a strong support to future developments in the forensic field [39]. The research involved discrimination among patients with stable angina, non- ST elevation myocardial infarction and AMI [38], and monitoring of atherosclerotic patients for myocardial infarction prevention [66] (all them developed by LC-QTOF MS/MS or LC–MS QqQ).
Search for cardiovascular disease omics biomarkers using no traditional samples. The presence of C-reactive protein (CRP) molecules in saliva provides an opportunity for development of non-invasive assessments of cardiovascular diseases risk factors [67]. The use of this simple no invasive sample requires the establishment of salivary CRP reference ranges and their correlation with serum concentrations, which have not been investigated in detail yet. A subsequent step could be to study the metabolism of CRP and establish a relationship between the levels of some of its characteristic metabolites and the given disease. Another potential metabolomics biomarker research could result after the characteristics of dry sweat as compared to fresh sweat are well established. This sample could be used for forensic purposes in dealing with lung cancer screening [10].
The key appearance of metabolites from wound-healing processes. A number of wound healing-related molecules (e.g., IL-1b, IL-6, TNF-a, IFN-g, MCP-1, CXCL12, VEGF-A, EGF, KGF, pro-col Ia2 and pro-col IIIa1, and also fibroblasts and fibrocyte spartially transformed into contractile myofibroblasts) have provided fundamental information for wound age estimation by comprehensive analysis of these molecules from the perspective of morphometrics, protein and gene expressions. This information can be of great interest in forensic analysis [9]. Transformations of these molecules during the wound healing process undoubtedly produce characteristic metabolites that can provide information on the process by simpler, cheaper and faster analyses. Nevertheless, no metabolomics biomarkers of wound-healing processes have been reported so far.
New drugs, new metabolites, new analytical methods for them. The endless appearance of new design drugs of abuse makes mandatory the continuous research on methods for their identification and that of their metabolites, crucial for the assessment of drug exposure [68]. In general, these metabolites are formed only in vivo and are not available as analogs on the illicit drug market. Their presence in biological samples such as urine and hair positively confirms exposure to the parent drug. Also, analysis of drug metabolites reduces the likelihood of false negatives when the parent drug is not detected [69] as the metabolites of many drugs of abuse can be detected in urine for days and sometimes weeks after exposure [70].
The effect of weathering on biomarkers. As in other biomarker fields forensic metabolomics biomarkers require to study the effects of weathering on the given metabolites as a function of the climatic conditions to which the target has been subjected [7].
Use of diseases-associated biomarkers. New proposals on biomarkers in the forensics field include the use of diseases-associated biomarkers as a provisional tool for identifying individuals in complicated forensic challenges. Such is the case of systemic lupus erythematosis, one among the diseases associated with the release of different biomarkers in body fluids. These biomarkers could help a differential identification of individuals at acceptable level of validity; however, the main drawback to face this proposal is the previous establishment of a disease-related biomarker bank [71].
Better knowledge about the metabolism of potential biomarkers. An in-depth study on the metabolism of biomarkers based on any omics is mandatory for accurate monitoring of a given compound through the alternative metabolic pathways it can follow depending on personal characteristics of the target individual, thus avoiding or reducing the possibility of false negatives.
Evolution of forensic psychiatric diagnosis toward omics biomarkers. The recent publication by Brown “From bibles to biomarkers: the future of the statistical manual on mental disorders (DSM) and forensic psychiatric diagnosis”shows as the once immovable psychiatric world has evolved towards the use of omics biomarkers that may involve metabolomics [72].
Acknowledgements
Funding by the Andalusian Regional Government (Junta de Andalucía) and FEDER program through project FQM-1602 is gratefully acknowledged.
References
- Eganhouse RP. Molecular markers in environmental geochemistry. Am Chem Soc Symp, ACS, Washington DC;1997.
- Jain KK. The handbook of biomarkers. Humana Press, New York. 2010.
- Strimbu K, Tavel JA. What are biomarkers? Curr Opin HIV AIDS. 2010 Nov;5(6):463–466. Pubmed Central PMCID: PMC3078627.
- Medeiros PM, Simoneit BR. Gas chromatography coupled to mass spectrometry for analyses of organic compounds and biomarkers as tracers for geologica, environmental, and forensic research. J SepSci. 2007 Jul;30(10):1516–1536. PubMed PMId: 17623433.
- Yang C, Wang ZD, Hollebone B, Brown CE, Landriault M. Application of light petroleum biomarkers for forensic characterization and source identification of spilled light refined oils. Environ Forensics. 2012 Jan;13(4):298–311.
- Wang Z, Stout SA, Fingas M. Forensic fingerprinting of biomarkers oil spill characterization and source identification. Environ Forensics. 2006 Feb;7(2):105-146.
- Malmborg J. Examining the weathering susceptibility of light biomarkers. Environ Forensic. 2017 Aug;18(3):197–206.
- Schröck A, Hernández Redondo A, Marin Fabritius M, König S, Weinmann W. Phosphatidylethanol (PEth) in blood samples from “driving under the influence” cases as indicator for prolonged excessive alcohol consumption. Int J Legal Med. 2016 Mar;130(2):393–400. PubMed PMID: 26671597.
- Wang LL, Zhao R, Liu CS, Liu M, Li SS, Li JY, et al. A fundamental study on the dynamics of multiple biomarkers in mouse excisional wounds for wound age estimation. J Forensic Leg Med. 2016 Apr;39:138–146. PubMed PMID: 26889946.
- Fernández-Peralbo MD, Gómez-Gómez E, Calderón-Santiago M, Carrasco- Valiente J, Ruiz-García J, Requena-Tapia MJ, et al. Prostate cancer patientsnegative biopsy controls discrimination by untargeted metabolomics analysis of uine by LC–QTOF: Upstream information on other omics. Sci Rep. 6(38243):1-11.
- Calderón-Santiago M, Priego-Capote F, Turck N, Robin X, Jurado-Gámez B, Jurado-Gamez B, et al. Human sweat metabolomics for lung cancer screening. Anal Bioanal Chem. 2015 Jul;407(18):5381–5392. Pubmed PMID: 25935675.
- Benson S, Lennard C, Maynard P, Roux C. Forensic applications of isotope ratio mass spectrometry-A review. Forensic Sci Int. 2006 Feb;157(1):1–22. PubMed PMID: 15919168.
- Bonnie R, Wallace R. Elder mistreatment: abuse, neglect, and exploitation in an aging America. National research council (US) Panel to review risk and prevalence of elder abuse and neglect. National Academy Press, Washington DC; 2003.
- Pearsall C. Forensic biomarkers of elder abuse: what clinicians need to know. J Forensic Nurs. 2005;1(4):182–186. PubMed PMID: 17073122.
- Gill P, Jeffreys AJ, Werrett DJ. Forensic application of DNA “fingerprints”. Nature. 1985 Dec 18;318(6046):577–579. PubMed PMID: 3840867.
- Delgado-Povedano MM, Calderón-Santiago M, Priego-Capote F, Jurado- Gámez B, Luque de Castro MD. Recent advances in human sweat metabolomics for lung cáncer screening. Metabolomics. Nov 2016;12(11):166.
- Dumache R, Ciocan V, Muresan C, Enache A. Molecular genetics and its applications in forensic sciences. In: Shetty BSK, Padubidri JR, editors. Forensic analysis – from death to justice. InTech, London: UnitedKingdom.
- Mannucci A, Sullivan KM, Ivanov PL, Gill P. Forensic application of a rapid and quantitative DNA sex test by amplification of the X–Y homologous gene amelogenin. Int J Leg Med. 1994;106(4):190–193. Pubmed PMID: 8038111.
- Andreasson H, Allen M. Rapid quantification and sex determination of forensic evidence materials. J Forensic Sci. 2003 Nov;48(6):1280–1287. PubMed PMID: 14640271.
- Keating B, Bansal AT, Walsh S, Millman J, Newman J. First all-in-one diagnostic tool for DNA intelligence: genome-wide inference of biogeographic ancestry, appearance, relatedness, and sex with the Identitas v1 Forensic Chip. Int J Legal Med. 2013 May;127(3):559–572. PubMed PMID: 23149900.
- Thomspon WC, Taroni F, Aitken CG. How the probability of a false positive affect the value of DNA evidence. J Forensic Sci. 2003 Jan;48(1):1–8. PubMed PMID: 12570198.
- An JH, Shin K-J, Yang WI, Lee HY. Body fluid identification in forensics. BMB Rep. 2012 Oct;45(10):545–553. PubMed PMID: 23101507.
- Kayser M, de Knijff P. Improving human forensics through advances in genetics, genomics and molecular biology. Nat Rev Genet. 2011 Mar;12(3):179– 192. PubMed PMID: 21331090.
- Credle JJ, Itoh CY, Yuan T, Sharma R, Scott ER, Fan Y, et al. Multiplexed analysis of fixed tissue RNA using Ligation in situ Hybridization. Nucleic Acids Res. 2017 Aug 21; 45(14):1–9. PubMed PMID: 28854731.
- Kramer F, Halámková L, Poghossian A, Schoöning MJ, Katz E, Halamek J. Biocatalytic analysis of biomarkers for forensic identification of ethnicity between Caucasian and African American groups. Analyst. 2013 Nov 7;138(21):6251–6257. PubMed PMID: 24003440.
- James P. Protein identification in the post-genome era: the rapid rise of proteomics. Q Rev Biophys. 1997 Nov;30(4):279–331. PubMed PMID: 9634650.
- Bakshi S, Halámková L, Halamek J, Katz E. Biocatalytic analysis of biomarkers for forensic identification of gender. Analyst. 2014;139(3): 559–563.
- Katz E, Halamek J. New approach in forensic analysis-biomolecular computing based analysis of significant forensic biomarkers. Ann Forensic Res Anal. 2014 Mar 04;1(1): 1002–1005.
- Brunelle E, Halamek J. Biocomputing approach in forensic analysis. IJPEDS. 2017;32(1): 17–29.
- Agudelo J, Huynh C, Halamek J. Forensic determination of blood sample age using a bioaffinity-based assay. Analyst. 2015 Mar 7;140(5):1411-1415. PubMed PMID: 25612932.
- Amoresano A, Flagiello A, Balsamo M, Carpentieri A, Pucci P. Proteomics in forensic science. Rendiconti Accademia Nazionale delle Scienze delta dei XL. Memorie di Scienze Fisiche e Naturali. 77-86.
- Gaensslen RE. Sourcebook in forensic serology, immunology, and biochemistry. US Department of Justice; 1983 Aug.
- Elkins KM. Forensic DNA biology: a laboratory manual. 1st ed. Academic Press: Oxford; 2012 Sep. 224.
- Bauer M. RNA in forensic science. Forensic Sci Int Genet. 2007 Mar;1(1):69–74. PubMed PMID: 19083730.
- George R, Donald PM, Nagraj SK, Idiculla JJ, Ismail RH. The impact of chimerism in DNA-based forensic sex determination analysis. MJMS. 2013 Mar;20(1):75–79. PubMed Central PMCID: PMC3684380.
- Boccard J, Badoud F, Grata E, Ouertani S, Hanafi M, Lanteri P, et al. A steroidomic approach for biomarkers discovery in doping control. Forensic Sci Int. 2011 Dec 10;213(1-3):85–94. PubMed PMID: 21831550.
- Wren ME, Shirtcliff EA, Drury SS. Not all biofluids are created equal: Chewing over salivary diagnostics and the epigenome. Clin Ther. 2015 Mar 1;37(3):529–539. PubMed PMID: 25778408.
- Calderón-Santiago M, Priego-Capote F, Galache-Osuna JG, Luque de Castro MD. Metabolomic discrimination between patients with stable angina, non-ST elevation myocardial infarction, and acute myocardial infarct. Electroforesis. 2013 Oct;34(19): 2827–2835.
- Fernández-Peralbo MA, Priego-Capote F, Galache-Osuna JG, Luque de Castro MD. Targeted analysis of omega-6-derivatives of eicosanoids in human serum by SPE–LC–MS/MS for evaluation of coronary artery disease. Electrophoresis. 2013 Oct;34(19):2901–2009. PubMed PMID: 24228265.
- Rashaid AH. Clinical and forensic biomarkers in human hair. Ohio University, Ohio, United States; 2014.
- Rainville PD, Langridge JI, Wrona MD, Wilson ID, Plumb RS. Integration of microfluidic LC with HRMS for the analysis of analytes in biofluids: past, present and future. Bioanalysis. 2015;7(11):1397–1411. PubMed PMID: 26110713.
- Delgado-Povedano MM, Calderón-Santiago M, Priego-Capote F, Luque de Castro MD. Study of sample preparation for quantitative analysis of amino acids in human sweat by liquid chromatography–tandem mass spectrometry. Talanta. 2016;146:310–317. PubMed PMID: 26695269.
- Mena-Bravo A, Priego-Capote F, Luque de Castro MD. Study of blood collection and sample preparation for analysis of vitamin D and its metabolites by liquid chromatography–tandem mass spectrometry. Anal Chim Acta. 2015 Jun 16;879:69–76. PubMed PMID: 26002479.
- Thakare R, Chhonker YS, Gautam N, Abdullah Alamoudi J, Alnouti Y. Quantitative analysis of endogenous compounds. J Pharm Biomed Anal. 2016 Sep 5;128:426–437. PubMed PMID: 27344632.
- Huynh C, Brunelle E, Halámková L, Agudelo J, Halámek J. Forensic identification of gender from fingerprints. Anal Chem. 2015;87(22):11531– 11536.
- Mena-Bravo A, Luque de Castro MD. Sweat: a sample with limited present applications and promising future in metabolomics. J Pharm Biomed Anal. 2014 Mar;90:139–147. PubMed PMID: 24378610.
- Croxton RS, Baron MG, Butler D, Kent T, Sears VG. Variation in amino acid and lipid composition of latent fingerprints. Forensic Sci Int. 2010 Jun 15;199(1-3):93–102. PubMed PMID: 20413233.
- Meenu S, Sayanti S, Mayank B, Pragzna Y, Bommi D. Non-invasive diagnostic tool for pathological conditions: salivary biomarkers. Int J Pharm Biol Arch. 2014 May;5(3):1-12.
- Wilson AD. Electronic-nose applications in forensic science and for analysis of volatile biomarkers in the human breath. J Forensic Sci Criminol. 2014 Jun;1(1):1-21.
- Oppolzer D, Barroso M, Gallardo. Bioanalytical procedures and developments in the determination of alcohol biomarkers in biological specimens. Bioanalysis. 2016 Feb;8(3): 220–251. PubMed PMID: 26795230.
- Alladio E, Martyna A, Salomone A, Pirro V, Vincenti M, Zadora G. Direct and indirect alcohol biomarkers data collected in hair samples – multivariate data analysis and likelihood ratio interpretation perspectives. Data in Brief. 2017 Jun;12:1–8. PubMed Central PMCID: 5457474.
- Pirro V, Valente V, Oliveri P, De Bernardis A, Salomone A, Vincenti M. Chemometric evaluation of nine alcohol biomarkers in a large population of clinically-classified subjects: pre-eminence of ethyl glucuronide concentration in hair for confirmatory classification. Anal Bioanal Chem. 2011 Oct;401(7):2153–2164. PubMed PMID: 2190146.
- Kristoffersen L, Stormyhr LE, Smith-Kielland A. Headspace gas chromatographic determination of ethanol: the use of factorial design to study effects of blood storage and headspace conditions on ethanol stability and acetaldehyde formation in whole blood and plasma. Forensic Sci Int. 2006 Sep;161(1–2):151–157. PubMed PMID: 16843627.
- Varga A, Hansson P, Johnson G, Alling C. Normalization rate and cellular localization of phosphatidylethanol in whole blood from chronic alcoholics. Clin Chim Acta. 2000 Sep;299(1-2):141-150. PubMed PMID: 10900300.
- Mauck CK, Doncel GF. Biomarkers of semen in the vagina: applications in clinical trials of contraception and prevention of sexually transmitted pathogens including HIV. Contraception. 2007 Jun;75(6):407–419. PubMed PMID: 17519146.
- Jordan KW, Chen LL. NMR-based metabolomics approach to target biomarkers for human prostate cancer. Expert Rev Proteomics. 2007 Jun;4(3):389-400. PubMed PMID: 17552923.
- Kovac JR, Pastuszak AW, Lamb DJ. The use of genomics, proteomics, and metabolomics in identifying biomarkers of male infertility. Fertil Steril. 2013 Mar;99(4):998–1007. PubMed PMID: 23415969.
- Mangos JA, Donnelly WH. Isolated parotid acinar cells from patients with cystic fibrosis. Morphology and composition. J Dent Res. 1981 Jan;60(1):19–25. PubMed PMID: 6934186.
- Boat TF, Weisman UN, Pallavicini JC. Purification and properties of the calcium precipitable protein in submaxillary saliva of normal and cystic fibrosis subjects. Pediatr Res. 1974 May;8(5):531–534. PubMed PMID: 4464870.
- Wotman S, Mercadante J, Mandel ID, Goldman R, Denning CR. The occurrence of calculus in normal children, children with cystic fibrosis and children with asthma. J Periodontol. 1973 May;44(5):278–280. PubMedPMID: 4512218.
- Slomiany BL, Aono M, Murty VL, Slomiany A, Levine MJ, Tabak LA. Lipid composition of submandibular saliva from normal and cystic fibrosis individuals. J Dent Res. 1982 Oct; 61(10):1163–1166. PubMed PMID: 6956597.
- Mandel ID, Kutscher A, Denning CR, Thompson RH. Salivary studies in cystic fibrosis. Am J Dis Child. 1967;113(4):431–438. PubMed PMID: 502166.
- Xu F, Teng X, Yuan X, Sun J, Wu H, Tang Y, et al. LCK: a new biomarker candidate for the early diagnosis of acute myocardial infarction. Mol Biol Rep. 25014 Dec;41(12): 8047–8053. PubMed PMID: 25209966.
- Aydin S, Aydin S, Kobat MA, Kalayci M, Eren MN, Yilmaz M, et al. Decreased saliva/serum irisin concentrations in the acute myocardial infarction promising for being a new candidate biomarker for diagnosis of this pathology. Peptides. 2014 Jun;56:141–145.
- Antzelevitch C, Brugada P, Borggrefe M, Brugada J, Brugada R, et al. Brugada syndrome: report of the second consensus conference: endorsed by the Heart Rhythm Society and the European Heart Rhythm Association. Circulation. 2005 Feb 8;111(5):659–670. PubMed PMID: 15655131.
- Calderón-Santiago. Metabolomic markers to monitor atherosclerotic patients for myocardial infarction prevention. Pers comm.
- Dillon MC, Opris DC, Kopanczyk R, Licklter J, Cornwell HN, Nazar AM, et al. Detection of homocysteine and C-reactive protein in the saliva of healthy adults: comparison with blood levels. Biomark Insights. 2010 Jul 20;5:57–61. PubMed PMID: 20703322.
- Carmo H, Hengstler JG, de Boer D, Ringel M, Remiao F, Oesch F, et al. Metabolic pathways of 4-bromo-2,5-dimethoxyphenethylamine (2C-B): analysis of phase I metabolism with hepatocytes of six species including human. Toxicology. 2005 Jan 5;206(1):75–89. PubMed PMID: 15590110.
- Kanamori T, Kuwayama K, Tsujikawa K, Miyaguchi H, Iwata YT, Inoue H. Synthesis and identification of urinary metabolites of 4-iodo-2,5-dimethoxyphenethylamine.J Forensic Sci. 2011 Sep;56(5):1319–1323. PubMed PMID: 21595694.
- Meyer MR, Maurer HH. Metabolism of designer drugs of abuse: an updated review. Curr Drug Metab. 2010 Jun 1;11:468–482. PubMed PMID: 20540700.
- Eid HA. The use of systemic lupus erythematosis (SLE) biomarkers in forensic investigation: a suggested approach. J Forensic Res. 2014 Apr 5;S:S12- 003.
- Brown TR. From Bibles to Biomarkers: The future of the DSM and forensic psychiatric diagnosis. Utah L Rev. 2015;4:743–766.